|
 Data availability
All sequencing data have been publicly deposited in the Genome Sequence Archive database (https://ngdc.cncb.ac.cn/gsa) under the accession number of PRJCA041520. All data in this study are available from the corresponding author on request. Source data are provided with this paper.
Code availability
The code used to process the data is publicly available at GitHub (https://github.com/averycheng-Hulab/iCDC).
References
Conrad, N. et al. Temporal trends and patterns in heart failure incidence: a population-based study of 4 million individuals. Lancet 391, 572–580 (2018).
Article
PubMed
Google Scholar
Yan, T. et al. Data availability
All sequencing data have been publicly deposited in the Genome Sequence Archive database (https://ngdc.cncb.ac.cn/gsa) under the accession number of PRJCA041520. All data in this study are available from the corresponding author on request. Source data are provided with this paper.
Code availability
The code used to process the data is publicly available at GitHub (https://github.com/averycheng-Hulab/iCDC).
References
Conrad, N. et al. Temporal trends and patterns in heart failure incidence: a population-based study of 4 million individuals. Lancet 391, 572–580 (2018).
Article
PubMed
Google Scholar
Yan, T. et al.
Burden, trends, and inequalities of heart failure globally, 1990 to 2019: a secondary analysis based on the global burden of disease 2019 study. J. Am. Heart Assoc. 12, e027852 (2023).
Article
PubMed
PubMed Central
Google Scholar
Khan, M. S. et al. Global epidemiology of heart failure. Nat. Rev. Cardiol. 21, 717–734 (2024).
Article
PubMed
Google Scholar
Greene, S. J. et al. Worsening heart failure: nomenclature, epidemiology, and future directions: JACC review topic of the week. J. Am. Coll. Cardiol. 81, 413–424 (2023).
Article
PubMed
Google Scholar
Adamo, L., Rocha-Resende, C., Prabhu, S. D.  & Mann, D. L. Reappraising the role of inflammation in heart failure. Nat. Rev. Cardiol. 17, 269–285 (2020).
Article
PubMed
Google Scholar
Markousis-Mavrogenis, G. et al. Immunomodulation and immunopharmacology in heart failure. Nat. Rev. Cardiol. 21, 119–149 (2024).
Article
PubMed
Google Scholar
Sabado, R. L., Balan, S. & Bhardwaj, N. Dendritic cell-based immunotherapy. Cell Res. 27, 74–95 (2017).
Article
CAS
PubMed
Google Scholar
Morante-Palacios, O.,   Fondelli, F., Fondelli, F.,
Ballestar, E. & Martinez-Caceres, E. M. Tolerogenic dendritic cells in autoimmunity and inflammatory diseases. Trends Immunol. 42, 59–75 (2021).
Article
CAS
PubMed
Google Scholar
Virani, S. S. et al. Heart disease and stroke statistics—2021 update: a report from the American Heart Association. Circulation 143, e254–e743 (2021).
Article
PubMed
PubMed Central
Google Scholar
Ambrosy, A. P. et al. The global health and economic burden of hospitalizations for heart failure: lessons learned from hospitalized heart failure registries. J. Am. Coll. Cardiol. 63, 1123–1133 (2014).
Article
PubMed
Google Scholar
Jones, N. R., Roalfe, A.  K., Adoki, I., Hobbs, F. D. R. & Taylor, C. J. Survival of patients with chronic heart failure in the community: a systematic review and meta-analysis. Eur. J. Heart Fail. 21, 1306–1325 (2019).
Article
PubMed
PubMed Central
Google Scholar
Cohn, J. N.,   Ferrari, R. Ferrari, R.
& Sharpe, N. Cardiac remodeling—concepts and clinical implications: a consensus paper from an international forum on cardiac remodeling. J. Am. Coll. Cardiol. 35, 569–582 (2000).
Article
CAS
PubMed
Google Scholar
Frantz, S., Hundertmark, M. J., Schulz-Menger, J., Bengel, F. M. & Bauersachs, J. Left ventricular remodelling post-myocardial infarction: pathophysiology, imaging, and novel therapies. Eur. Heart J. 43, 2549–2561 (2022).
Article
CAS
PubMed
PubMed Central
Google Scholar
Mann, D. L. The emerging role of innate immunity in the heart and vascular system: for whom the cell tolls. Circ. Res.  108, 1133–1145 (2011).
Article
CAS
PubMed
PubMed Central
Google Scholar
Murphy, S. P., Kakkar, R., McCarthy, C. P. & Januzzi, J. L. Jr. Inflammation in heart failure: JACC state-of-the-art review. J. Am. Coll. Cardiol. 75, 1324–1340 (2020).
Article
PubMed
Google Scholar
Monaco, C. et al. Immunotherapy for atherosclerosis. Physiol. Rev.   105, 2141–2230 (2025).
Hackstein, H. 105, 2141–2230 (2025).
Hackstein, H.
& Thomson, A. W. Dendritic cells: emerging pharmacological targets of immunosuppressive drugs. Nat. Rev. Immunol. 4, 24–34 (2004).
Article
CAS
PubMed
Google Scholar
Tureci, O. et al. Cascades of transcriptional induction during dendritic cell maturation revealed by genome-wide expression analysis. FASEB J. 17, 836–847 (2003).
Article
CAS
PubMed
Google Scholar
Rurik, J. G. et al. CAR T cells produced in vivo to treat cardiac injury. Science 375, 91–96 (2022).
Article
ADS
CAS
PubMed
PubMed Central
Google Scholar
Aghajanian, H. et al. Targeting cardiac fibrosis with engineered T cells. Nature 573, 430–433 (2019).
Article
ADS
CAS
PubMed
PubMed Central
Google Scholar
Du, H. et al. CAR macrophages engineered in vivo for attenuating myocardial ischemia-reperfusion injury. Circ. Res. 137, 846–859 (2025).
Article
CAS
PubMed
Google Scholar
Wang, J. et al. CAR-macrophage therapy alleviates myocardial ischemia-reperfusion injury. Circ. Res. 135, 1161–1174 (2024).
Article
ADS
CAS
PubMed
Google Scholar
Rong, Z. et al. An effective approach to prevent immune rejection of human ESC-derived allografts. Cell Stem Cell 14, 121–130 (2014).
Article
CAS
PubMed
PubMed Central
Google Scholar
Todorova, D. et al. hESC-derived immune suppressive dendritic cells induce immune tolerance of parental hESC-derived allografts. EBioMedicine 62, 103120 (2020).
Article
CAS
PubMed
PubMed Central
Google Scholar
Greenwald, R.  J., Freeman, G. J. & Sharpe, A. H. The B7 family revisited. Annu. Rev. Immunol. 23, 515–548 (2005).
Article
PubMed
Google Scholar
Sharma, P. et al. Immune checkpoint therapy-current perspectives and future directions. Cell 186, 1652–1669 (2023).
Article
ADS
CAS
PubMed
Google Scholar
Buchbinder, E. I. & Desai, A. CTLA-4 and PD-1 pathways: similarities, differences, and implications of their inhibition. Am. J. Clin. Oncol.   39, 98–106 (2016).
Article
CAS
PubMed
PubMed Central
Google Scholar
Balaji, S. et al. Interleukin-10-mediated regenerative postnatal tissue repair is dependent on regulation of hyaluronan metabolism via fibroblast-specific STAT3 signaling. FASEB J. 39, 98–106 (2016).
Article
CAS
PubMed
PubMed Central
Google Scholar
Balaji, S. et al. Interleukin-10-mediated regenerative postnatal tissue repair is dependent on regulation of hyaluronan metabolism via fibroblast-specific STAT3 signaling. FASEB J.
31, 868–881 (2017).
Article
CAS
PubMed
Google Scholar
Saraiva, M., Vieira, P. & O’Garra, A. Biology and therapeutic potential of interleukin-10. J. Exp. Med. 217, e20190418 (2020).
Forte, E., Furtado, M. B. & Rosenthal, N. The interstitium in cardiac repair: role of the immune-stromal cell interplay. Nat. Rev. Cardiol. 15, 601–616 (2018).
Article
CAS
PubMed
Google Scholar
Bhattacharya, M. & Ramachandran, P. Immunology of human fibrosis. Nat. Immunol.  24, 1423–1433 (2023).
Article
CAS
PubMed
Google Scholar
Kim, J. et al. Wild-type p53 promotes cancer metabolic switch by inducing PUMA-dependent suppression of oxidative phosphorylation. Cancer Cell 35, 191–203, (2019).
Article
CAS
PubMed
Google Scholar
Pinto, A. R. et al. Revisiting cardiac cellular composition. Circ. Res. 118, 400–409 (2016).
Article
CAS
PubMed
Google Scholar
Yao, L. et al. Spatial multiplexed protein profiling of cardiac ischemia-reperfusion injury. Circ. Res. 133, 86–103 (2023).
Article
CAS
PubMed
Google Scholar
Li, Q. et al. Small extracellular vesicles containing miR-486-5p promote angiogenesis after myocardial infarction in mice and nonhuman primates. Sci. Transl. Med. 13, eabb0202 (2021).
Lindsey, M. L. et al. Guidelines for experimental models of myocardial ischemia and infarction. Am. J. Physiol. Heart. Circ. Physiol. 314, H812–H838 (2018).
Article
CAS
PubMed
PubMed Central
Google Scholar
Lindsey, M. L. et al. Guidelines for in vivo mouse models of myocardial infarction. Am. J. Physiol. Heart. Circ. Physiol. 321, H1056–H1073 (2021).
Article
CAS
PubMed
PubMed Central
Google Scholar
Xiao, C. et al. FMO2 prevents pathological cardiac hypertrophy by maintaining the ER-mitochondria association through interaction with IP3R2-Grp75-VDAC1. Circulation 151, 1667–1685 (2025).
Article
CAS
PubMed
Google Scholar
Hu, X. et al. A large-scale investigation of hypoxia-preconditioned allogeneic mesenchymal stem cells for myocardial repair in nonhuman primates: paracrine activity without remuscularization. Circ. Res.   118, 970–983 (2016).
Article
ADS
CAS
PubMed
PubMed Central
Google Scholar
Li, G. et al. Dynamic molecular atlas of cardiac fibrosis at single-cell resolution shows CD248 in cardiac fibroblasts orchestrates interactions with immune cells. Nat Cardiovasc Res. 118, 970–983 (2016).
Article
ADS
CAS
PubMed
PubMed Central
Google Scholar
Li, G. et al. Dynamic molecular atlas of cardiac fibrosis at single-cell resolution shows CD248 in cardiac fibroblasts orchestrates interactions with immune cells. Nat Cardiovasc Res.
4, 380–396 (2025).
Article
CAS
PubMed
Google Scholar
Ackers-Johnson, M. et al. A simplified, Langendorff-free method for concomitant isolation of viable cardiac myocytes and nonmyocytes from the adult mouse heart. Circ. Res. 119, 909–920 (2016).
Article
CAS
PubMed
PubMed Central
Google Scholar
Download references
Acknowledgements
We thank Y. Wang for discussion; and T. Lin, Y. Li, T. Lu, Y. Wang and L. Wang for technical support. This work was supported by National Science Fund for Distinguished Young Scholars (no. 82225004 for X. Hu), grants from National Key Research and Development Program of China (no. 2023YFA1800700 for X. Hu and 2022YFC3401600 for Yang Xu), the Key Project of National Natural Science Foundation of China (no. 82430013 for X. Hu and 82230102 for Yang Xu), Leading Innovative and Entrepreneur Team Introduction Program of Zhejiang (no. 2022R01002 for Yang Xu), the Project for Central-Guided Regional Science and Technology Development (no. 2024ZY01044 for X. Hu), the Key research and development program of Zhejiang Province (no.  2022C03006 for Yang Xu) and the Fundamental Research Funds for the Central Universities (no. 2021FZZX001-42 for Yang Xu).
Author information
Author notes
These authors contributed equally: Xiaoying Li, Jiamin Li, Guohua Li, Lisheng Zhu, Guo Cheng, Huanqiang Li
Authors and Affiliations
Department of Cardiology, The Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China
Xiaoying Li, Jiamin Li, Guohua Li, Guo Cheng, Huanqiang Li, Hao Lin, Ningqing Jia, Xiaoqian Hong, Ye Liu, Zhiwei Zhong, Yize Chen, Biqing Wang, Jing Zhao, Zhenqi Hua, Lingjun Wang, Qiming Chen, Peijie Zheng, Shuyuan Sheng, Songting Gu, Cheng Ni, Shuchang Ye, Changle Ke, Feimu Zhang, Mo Li, Junhua He, Yan Wu, Yinghui Xu, Huajun Li, Yu Zhang, Guanhua Hu, Chengchen Zhao, Yang Xu & Xinyang Hu
State Key Laboratory of Transvascular Implantation Devices, Hangzhou, China
Xiaoying Li, Guohua Li, Huanqiang Li, Hao Lin, Ningqing Jia, Xiaoqian Hong, Jing Zhao, Zhenqi Hua, Qiming Chen, Peijie Zheng, Yan Wu, Yang Xu & Xinyang Hu
Heart Regeneration and Repair Key Laboratory of Zhejiang Province, Hangzhou, China
Xiaoying Li, Guohua Li, Huanqiang Li, Hao Lin, Ningqing Jia, Xiaoqian Hong, Jing Zhao, Zhenqi Hua, Qiming Chen, Peijie Zheng, Yan Wu, Yang Xu & Xinyang Hu
Binjiang Institute of Zhejiang University, Hangzhou, China
Xiaoying Li, Guo Cheng, Lingjun Wang, Yang Xu & Xinyang Hu
Transvascular Implantation Devices Research Institute, Hangzhou, China
Jiamin Li & Xinyang Hu
Department of Medical Oncology, The Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China
Lisheng Zhu & Lili Yu
Liangzhu Laboratory, Zhejiang University, Hangzhou, China
Shaohui Shi
Department of Cardiothoracic Surgery, The Second Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, China
Minjian Kong
Department of Anesthesiology, The Second Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, China
Qi Chen
Department of Radiology, The Second Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, China
Jianzhong Sun
Department of Ultrasound in Medicine, The Second Affliated Hospital, Zhejiang University School of Medicine, Hangzhou, China
Yiping Dong
Contributions
X.L., J.L., G.L., Huanqiang Li, H. Lin, L.Z., N.J., X. Hong, Z.H.,   L.W., Qiming Chen, P.Z., S. L.W., Qiming Chen, P.Z., S.
Sheng, Y.C., S.G., Y.Z., G.H. and L.Y. performed mouse surgeries and subsequent downstream experiments. X.L., L.Z. and Huanqiang Li generated the lentiviral construct and lentivirus, and engineered DCs and conducted in vitro assays. M.K., Qi Chen and C.K. established the MI model in NHPs. Huajun Li and Y.D. performed cardiac echocardiography assessments in NHPs. J.S. and C.Z. conducted MRI analyses in NHPs.  J.L., Huanqiang Li, J.Z. Y.C. and L.Z. carried out western blot experiments. J.L., H. Lin and S.Y. performed Sirius Red staining and image analysis. J.L., H. Lin, Z.Z., S. Sheng, S.Y., F.Z. and J.H. conducted immunofluorescence staining and data quantification. G.C., M.L. and S. Shi performed scRNA-seq and spatial transcriptomic analyses.   Y.L., Y.L.,
B.W., Y.W. and Yinghui Xu organized and analysed the data. X.L., J.L., G.L., Huanqiang Li and C.N. interpreted the data and contributed to experimental design and manuscript revision. Yand Xu and X. Hu conceived the study, supervised the project, interpreted the data, and drafted and edited the manuscript. All of the authors read and approved the final version of the manuscript.
Corresponding authors
Correspondence to
Yang Xu or Xinyang Hu.
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Betul Gunes, Kevin King, Louai Labanieh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
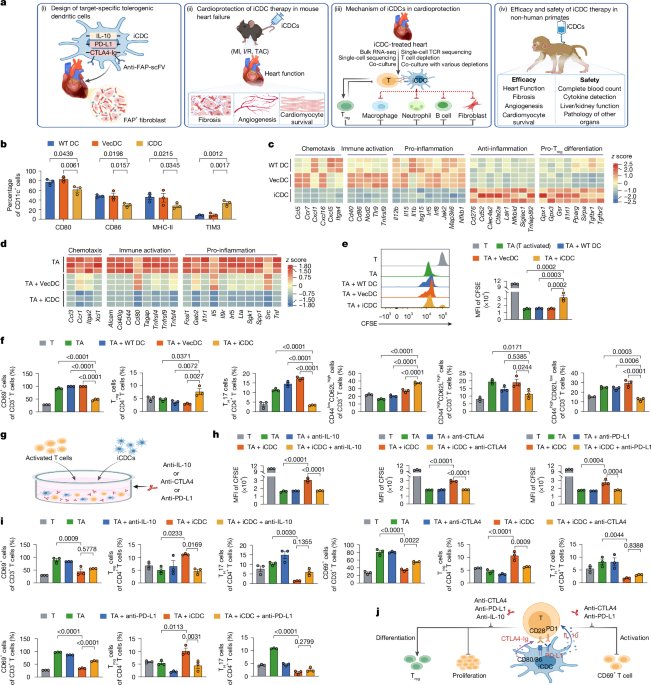
Extended data figures and tablesExtended Data Fig. 1 Generation and in vitro characterization of FAP-targeted immunosuppressive dendritic cells (iCDCs).
a, Schematic of the lentiviral construct encoding a fibroblast activation protein (FAP)-specific chimeric antigen receptor (CAR) composed of an anti-FAP scFv, murine CD8α hinge/transmembrane domains, and IRES-linked immunomodulatory payloads: CTLA4-Ig, PD-L1, and IL-10.  b, Representative flow cytometry plots showing CAR expression in cultured CD11c+ dendritic cells (DCs) transduced with wild-type (WTDC), vector control (VecDC), or immunosuppressive (iCDC) constructs. c, ELISA quantification of IL-10 and CTLA4-Ig secretion from DC culture supernatants, showing significantly elevated production in iCDCs compared to WTDC and VecDC. n = 3 biological replicates per group. d, Flow cytometry analysis of co-stimulatory molecules (CD80, CD86, MHC-II) and inhibitory immune checkpoints (PD-L1, TIM3) in cultured CD11c+ DCs. iCDCs exhibited upregulation of TIM3 and PD-L1 as well as downregulation of CD80/CD86 and MHC-II. e,f, Gene Ontology (GO) and KEGG pathway enrichment analyses of differentially expressed genes comparing iCDCs with WTDCs (e) and VecDCs (f), highlighting the downregulation of inflammatory and immune activation pathways. Heatmaps display representative immunoregulatory gene expression (Z-score scaled). g–h, GO/KEGG pathway analysis of CD3/CD28-activated T cells cocultured with iCDCs (TA+iCDC) versus those cocultured with VecDCs (TA+VecDC, g) and uncocultured controls (TA, h), revealing broad suppression of chemotaxis, immune activation, and pro-inflammation pathways. i, Flow cytometric analysis of naive (CD62L+CD44−), central memory (TCM: CD62L+CD44+), and effector memory (TEM: CD62L−CD44+) T cell subsets in in vitro CD3/CD28-activated T cells cocultured with or without iCDCs in the presence or absence of neutralizing antibodies against IL-10, CTLA4, or PD-L1 (n = 3 biological replicates). Data are presented as mean ± s.e.m.; hypergeometric test (e–h), one-way ANOVA with Tukey’s multiple-comparisons test (c, i); the statistical tests other than e–h are two-sided.
Source data
Extended Data Fig.   2 Temporal dynamics of FAP expression and optimization of iCDC dosing and timing after ischaemia/reperfusion injury.
a, Western blot analysis of FAP protein expression in cardiac tissue at days 1, 3, 7, 14, and 28 post-MI. 2 Temporal dynamics of FAP expression and optimization of iCDC dosing and timing after ischaemia/reperfusion injury.
a, Western blot analysis of FAP protein expression in cardiac tissue at days 1, 3, 7, 14, and 28 post-MI.
b, Quantitative RT-PCR analysis of Fap mRNA level in heart and major organs at baseline and indicated time points post-MI, normalized to sham-operated hearts. c, Western blot analysis of FAP protein expression in hearts at day 1, 3, 7, 14, and 28 post-I/R. n = 6 mice per group in a–c. d, Immunofluorescence staining demonstrating co-localization of FAP (green) with vimentin+ fibroblasts (red) in peri-infarct regions at days 3 and 7 post-I/R. Nuclei, DAPI (blue). Scale bars, 50 μm. e, f, Representative M-mode echocardiograms and quantification of EF%, FS%, LVIDd and LVIDs at days 1 and 28 after I/R in mice treated with iCDCs on day 1 (D1), day 3 (D3) or day 7 (D7). n = 8 mice in iCDC (D1) and iCDC (D3), n = 9 mice in other three groups.  g, Sirius Red staining and fibrosis quantification at day 28 post-I/R. n = 6 mice per group. Scale bar, 2 mm. h–j, Dose-response analysis comparing low-dose (1.5 × 106) and high-dose (3.0 × 106) iCDC therapy after I/R. Representative echocardiograms, functional quantification and fibrosis analysis are shown. Sham, n = 7 mice; I/R, n = 11 mice; iCDC Low, n = 10 mice; iCDC High, n = 11 mice; fibrosis, n = 6 mice per group. Scale bar, 2 mm. k, l, Flow cytometry and ELISA validation of engineered DC phenotypes and IL-10/CTLA4-Ig secretion in culture (n = 3 biological replicates).   m–o, Ex vivo fluorescence imaging of DiR-labelled DCs with quantification (n = 3 mice per group). m–o, Ex vivo fluorescence imaging of DiR-labelled DCs with quantification (n = 3 mice per group).
p, Flow cytometry validation of FAP scFv expression in cultured WTDC, VecDC, CPIDC, and iCDC. q, r, Representative M-mode echocardiograms and quantification of LVIDd and LVIDs at day 3 and day 28 post-MI. n = 10 mice in Sham group, and n = 15 mice in other four groups. s, Kaplan-Meier survival curves for MI mice treated with VecDC or iCDC. n = 29 mice in MI group, and n = 25 mice in VecDC and iCDC groups. t, u, Representative M-mode echocardiograms and quantification of LVIDd and LVIDs at 3 days, 4 weeks, 8 weeks, and 12 weeks post-I/R. Sham, n = 8 mice; I/R, n = 11 mice; VecDC, n = 10 mice; iCDC, n = 10 mice.  v, w, Immunofluorescence (n = 6 mice per group) and transcriptomic analyses showing enhanced angiogenesis in iCDC-treated hearts. Data are shown as mean ± s.e.m. (b, f, i, l, n, q) or mean ± s.d. (a, c, g, j, t, u). All statistical tests are two-sided; one-way ANOVA with Tukey’s multiple-comparisons test (a, c, g, j, l, u), two-way ANOVA with Tukey’s multiple-comparisons test (b, f, i, q, t), and Kaplan–Meier survival analysis (r). Source data for immunoblots are provided in Supplementary Fig. 1.
Source data
Extended Data Fig. 3 Functional dissection of immunomodulatory payloads in iCDCs.
a, Flow cytometric validation of G4S linker expression in dendritic cells engineered to express dual immunomodulatory payloads together with FAP scFv, including FAP scFv+CTLA4-Ig+PD-L1 (FAP–CP), FAP scFv+IL-10 + CTLA4-Ig (FAP-CI), FAP scFv+IL-10 + PD-L1 (FAP-PI), and fully engineered iCDCs. b, c, Representative M-mode echocardiogram images and quantitative assessment of cardiac function at days 3 and 28 post-MI, including EF%, FS%, LVIDd and LVIDs. n = 10 mice in Sham group, and n = 15 mice in other five groups. d, Sirius Red staining and quantification of fibrotic area at day 28 post-MI in Sham (n = 6 mice), MI (n = 9 mice), FAP-CP (n = 8 mice), FAP-CI (n = 9 mice), FAP-PI (n = 8 mice), and iCDC (n = 10 mice) groups. Scale bar, 2 mm.   e, Western-blot analysis of fibronectin, periostin, and α-SMA in heart tissues with quantification normalized to β-actin (n = 6 mice per group). e, Western-blot analysis of fibronectin, periostin, and α-SMA in heart tissues with quantification normalized to β-actin (n = 6 mice per group).
f, Flow cytometry validation of FAP scFv expression in cultured dendritic cells expressing single immunomodulatory factor retaining FAP scFv, including FAP scFv+CTLA4-Ig (FAP-C), FAP scFv+PD-L1 (FAP-P), FAP scFv+IL-10 (FAP-I) and iCDCs. g, h, Representative M-mode echocardiograms and functional quantification at days 3 and 28 post-MI. n = 10 mice in Sham group, and n = 15 mice in other five groups. i, Sirius Red staining and fibrosis quantification at 4 weeks post-MI in Sham (n = 6 mice), MI (n = 8 mice), FAP-C (n = 8 mice), FAP-P (n = 8 mice), FAP-I (n = 7 mice), and iCDC (n = 8 mice) groups. Scale bar, 2 mm. j, Western-blot analysis of fibronectin, periostin, and α-SMA in heart tissues from Sham, MI, FAP-C, FAP-P, FAP-I, and iCDC groups, with quantification normalized to β-actin (n = 6 mice per group). Data are presented as mean ± s.e.m., except for d and j, shown as mean ± s.d. All statistical tests are two-sided; one-way ANOVA with Tukey’s multiple-comparisons test (d, e, i, j), and two-way ANOVA with Tukey’s multiple-comparisons test (c, h). Source data for immunoblots are provided in Supplementary Fig. 1.
Source data
Extended Data Fig.  4 FAP-targeted iCDC therapy attenuates cardiac dysfunction and fibrosis in pressure overload-induced heart failure.
a, Ex vivo bioluminescence imaging of hearts after intravenous infusion of DiR-labelled iCDCs, vector-only DCs (VecDC), DCs expressing CTLA4-Ig, PD-L1 and IL-10 without FAP scFv (CPI-DCs), and DCs expressing GFP-specific scFv, CTLA4-Ig, PD-L1 and IL-10 (GFP-CPI-DCs) in the TAC model. Right, signal quantification (n = 3 mice per group). b, Experimental timeline showing intravenous administration of iCDCs at 4 and 6 weeks post-TAC, with echocardiographic and histological analyses at 4, 6 and 8 weeks. c, Echocardiographic assessment demonstrating improved ejection fraction (EF%) and fractional shortening (FS%) in iCDC-treated mice. Sham, n = 11 mice; TAC, n = 10 mice; TAC+VecDC, n = 9 mice; TAC+iCDC, n = 8 mice. d, Sirius Red staining of cardiac sections at 8 weeks post-TAC showing reduced interstitial fibrosis with iCDC treatment; right, fibrosis quantification. Sham (n = 11 mice), TAC (n = 9 mice), VecDC (n = 9 mice), iCDC (n = 8 mice). Scale bars, 1 mm (upper) and 50 μm (lower).   e, Western blot analysis of fibronectin, periostin and α-SMA in heart tissues with quantification normalized to β-actin (n = 6 mice). e, Western blot analysis of fibronectin, periostin and α-SMA in heart tissues with quantification normalized to β-actin (n = 6 mice).
f, Kaplan–Meier survival analysis of TAC mice. TAC and VecDC, n = 14 mice; iCDC, n = 15 mice. g, Schematic of therapeutic intervention in mice with established heart failure (baseline EF < 40%). h, Echocardiographic evaluation showing improved EF% and FS% at 2 and 4 weeks after iCDC treatment in mice with baseline EF < 40%. n = 8 mice in Sham, n = 10 mice in other three groups. i, Sirius Red staining and fibrosis quantification at 8 weeks post-TAC. n = 6 mice in iCDC, n = 5 mice in other three groups. Scale bars, 1 mm (upper) and 50 μm (lower). j, Western blot analysis of fibrotic markers in hearts from Sham, TAC, TAC+VecDC and TAC+iCDC groups (n = 6 mice per group).  k, Immunofluorescence analysis of CD31+ endothelial cells showing increased vessel density after iCDC treatment. n = 6 mice in iCDC, n = 5 mice in other three groups. Scale bar, 500 μm (upper) and 100 μm (lower). l, Kaplan–Meier survival curves of TAC mice with baseline EF < 40%. n = 11 mice in iCDC, n = 12 mice in other two groups. Data are presented as mean ± s.e.m., except for k shown as mean ± s.d.; all statistical tests are two-sided; two-way ANOVA with Tukey’s multiple-comparisons test (a, c, h), one-way ANOVA with Tukey’s multiple-comparisons test (d, e, i–k), and Kaplan-Meier survival analysis by the Mantel-Cox test (f, l). Source data for immunoblots are provided in Supplementary Fig. 1. b and g were created with BioRender.com.
Source data
Extended Data Fig. 5 FAP-targeted iCDC therapy improves cardiac function and promotes angiogenesis in the TAC model.
a, Western blot analysis of FAP protein expression in cardiac tissue at 2, 4, 6, and 8 weeks after transverse aortic constriction (TAC), with quantification shown (n = 4 mice). b–d, Ex vivo fluorescence imaging of major organs after infusion of DiR-labelled WTDC, vector-only DCs (VecDC), DCs expressing CTLA4-Ig, PD-L1 and IL-10 without FAP scFv (CPIDC), and DCs expressing GFP-specific scFv, CTLA4-Ig, PD-L1 and IL-10 (GFP-CPIDC), and iCDC at 1 day, 3 days, 7 days and 14 days post-injection, with quantification shown in d (n = 3 mice per group). e, Representative M-mode echocardiography images of mice at 8 weeks post-TAC in Sham, TAC, TAC+VecDC, and TAC+iCDC groups.   f, Quantification of left ventricular internal dimensions in diastole (LVIDd) and systole (LVIDs) at 4 weeks, 6 weeks, and 8 weeks post-TAC across the indicated groups. Sham (n = 11 mice), TAC (n = 10 mice), VecDC (n = 9 mice), iCDC (n = 8 mice). f, Quantification of left ventricular internal dimensions in diastole (LVIDd) and systole (LVIDs) at 4 weeks, 6 weeks, and 8 weeks post-TAC across the indicated groups. Sham (n = 11 mice), TAC (n = 10 mice), VecDC (n = 9 mice), iCDC (n = 8 mice).
g, Immunofluorescence staining of heart sections showing CD31+ endothelial cells (red) in the indicated treatment groups. iCDC treatment markedly enhances capillary density. Sham (n = 11 mice), TAC (n = 9 mice), VecDC (n = 9 mice), iCDC (n = 8 mice). Scale bars, 500 μm (upper panel) and 100 μm (lower panel). h, Representative M-mode echocardiography images from TAC-injured mice with established heart failure (EF ≤ 40%) at 4 weeks post-treatment. i, LVIDd and LVIDs measurements at baseline (animal inclusion), and 2 weeks and 4 weeks post-therapy in mice with pre-existing heart failure. n = 8 mice in Sham, n = 10 mice in other three groups. Data are presented as mean ± s.e.m., except for e, f and h shown as mean ± s.d.; all statistical tests are two-sided; one-way ANOVA with Tukey’s multiple-comparisons test (a, f), and two-way ANOVA with Tukey’s multiple-comparisons test (e, h). Source data for immunoblots are provided in Supplementary Fig. 1.
Source data
Extended Data Fig. 6 Systemic safety assessment of iCDC treatment in MI, I/R, and TAC models.
a, Quantification of non-sustained VT episodes and duration, sustained VT episodes and duration, and PVCs numbers in MI mice treated with or without iCDC (n = 8 mice per group). b, H&E staining of lung, kidney, and liver tissues from Sham, MI, VecDC-, CPI-DC, and iCDC-treated groups at 28 days post-MI.  Scale bar, 100 μm. c, Serum analysis of ALT, AST, BUN, and Cr levels in the MI model (n = 6 mice per group). d, Representative hematoxylin and eosin (H&E) staining of lung, kidney, and liver tissues from Sham, I/R, VecDC-, and iCDC-treated mice at 28 days post-I/R injury. Scale bar, 100 μm. e, Serum biochemical analysis of liver and kidney function markers including alanine aminotransferase (ALT), aspartate aminotransferase (AST), blood urea nitrogen (BUN), and creatinine (Cr). n = 5 mice in VecDC, n = 6 mice in other three groups. f, H&E staining of lung, kidney, and liver from Sham, TAC, VecDC-, and iCDC-treated mice at 28 days post-TAC surgery.   Scale bar, 100 μm. Scale bar, 100 μm.
g, Corresponding serum biochemical assessment (ALT, AST, BUN, Cr) in the TAC model shows comparable values across groups. Sham (n = 6 mice), TAC (n = 5 mice), VecDC (n = 6 mice), iCDC (n = 7 mice). h, Serum cytokine levels measured by ELISA, including IFN-γ, IL-6, TNF-α, IL-17A, IL-4, and IL-10. iCDC treatment significantly reduced pro-inflammatory cytokines (IL-6, TNF-α) compared to I/R controls (n = 5 mice per group). Data are presented as mean ± s.e.m., except for h shown as mean ± s.d.; all statistical tests are two-sided; unpaired two-tailed Student’s t-test (a) and one-way ANOVA with Tukey’s multiple-comparisons test (c, e, g, h).
Source data
Extended Data Fig. 7 iCDC treatment reshapes the cardiac transcriptional landscape following I/R injury.
a, Gene Ontology (GO) enrichment analysis of upregulated (upper panel) and downregulated (lower panel) biological processes in iCDC-treated hearts compared to VecDC-treated controls. iCDC n = 6 mice, VecDC n = 4 mice. b, KEGG enrichment analysis of upregulated (left panel) and downregulated (right panel) genes comparing iCDC versus VecDC-treated groups. These results indicate that iCDC treatment attenuates pro-inflammatory and fibrotic signalling while promoting pathways associated with cardiac structural and metabolic recovery. P values in a and b were calculated using a hypergeometric test.
Extended Data Fig. 8 Single-Cell RNA Sequencing and flow cytometry reveal iCDC-mediated reprogramming of cardiac T cells following I/R injury.
a, Dot plot showing expression of representative marker genes across major cardiac immune cell subsets, including macrophage, dendritic cells, neutrophils, B cells, T cells, and natural killer (NK) cells, and fibroblasts and endothelial cells.  b, Dot plot displaying top marker genes across identified T cell subtypes, including T helper (T-Helper), cytotoxic T (T-Toxic), γδ T cells (T-γδ), regulatory T cells (Treg), naive T cells, central memory T cells (T-CM), and immature T cells. c, GO and KEGG enrichment analysis of downregulated pathways in cardiac T cells from iCDC-treated versus I/R control mice, revealing the suppression of pathways associated with T cell proliferation and activation (n = 3 pooled samples from 9 mice per group). d, Flow cytometric analysis of cardiac T cells from Sham, I/R, VecDC, and iCDC-treated mice, assessing the frequency of total CD3+ T cells, activated T cells (CD69+), TH17 cells (CD4+IL-17A+), regulatory T cells (CD4+CD25+Foxp3+), naive T cells (CD44−CD62L+), and memory subsets including central memory (TCM, CD44+CD62L+) and effector memory (TEM, CD44+CD62L−) T cells (n = 5 mice per group). e, Longitudinal flow cytometry analysis of cardiac CD4+CD25+Foxp3+ Tregs at day 7 and day 28 post-MI, assessing CD69, CD44, and CD62L expression to evaluate activation and memory differentiation (n = 5 mice per group). f–h, Single-cell TCR sequencing of cardiac and circulating CD4+ T cells at day 7 post-MI. UMAP projection identifies seven transcriptionally distinct T-cell clusters; stacked bar plots show relative cluster proportions in heart and blood; dot plots display representative marker genes. i, j, GO enrichment analysis of upregulated biological processes in cardiac regulatory T (Treg) cells (i) and expanded Treg cells (j) from iCDC-treated versus MI control mice. P values in c, i and l were calculated using a hypergeometric test.
Extended Data Fig. 9 Single-Cell RNA Sequencing reveals iCDC-mediated transcriptional modulation of cardiac macrophages and dendritic cells following ischaemia/reperfusion injury.
a, Dot plot showing top marker genes across identified cardiac myeloid subsets, including the subpopulations of macrophage, monocytes, and dendritic cells (DCs).   b, Gene Ontology (GO) and KEGG pathway enrichment analysis of downregulated genes in macrophages following iCDC treatment versus I/R alone (n = 3 pooled samples from 9 mice). Pathways involved in phagocytosis, cytokine production, and antigen presentation were significantly suppressed. The heatmap shows expression of representative downregulated genes associated with phagocytosis, antigen processing and presentation, and pro-inflammation in macrophages across Sham, I/R, and iCDC groups. b, Gene Ontology (GO) and KEGG pathway enrichment analysis of downregulated genes in macrophages following iCDC treatment versus I/R alone (n = 3 pooled samples from 9 mice). Pathways involved in phagocytosis, cytokine production, and antigen presentation were significantly suppressed. The heatmap shows expression of representative downregulated genes associated with phagocytosis, antigen processing and presentation, and pro-inflammation in macrophages across Sham, I/R, and iCDC groups.
c, Flow cytometry analysis of cardiac macrophages across treatment groups (Sham, I/R, VecDC, iCDC), showing changes in the expression of macrophage markers CCR2, MHCII, and TIM4. d, GO and KEGG enrichment analysis of downregulated genes in dendritic cells from iCDC-treated versus I/R control hearts, highlighting reduced expression in pathways related to antigen processing, endocytosis, and inflammatory signalling (n = 3 pooled samples from 9 mice). Accompanying heatmap displays expression of representative suppressed genes associated with antigen processing and presentation, immune activation, and pro-inflammation. e, GO and KEGG enrichment analysis of upregulated genes in dendritic cells from iCDC-treated versus I/R control hearts (n = 3 pooled samples from 9 mice). Enriched pathways include those related to immune regulation, cell adhesion, and tissue repair. Heatmap at right illustrates increased expression of representative genes associated with cell adhesion, cell migration and anti-inflammation in iCDC-treated hearts. P values in b, d, and e were calculated using a hypergeometric test.
Extended Data Fig. 10 Single-Cell RNA Sequencing and flow cytometry analysis of neutrophils and B cells in response to iCDC treatment.
a, Dot plot showing expression of representative marker genes across cardiac neutrophil subpopulations. b, Gene Ontology (GO) and KEGG pathway enrichment analysis of genes downregulated in cardiac neutrophils from iCDC-treated versus I/R control hearts, revealing inhibition of inflammatory responses, LPS sensing, and neutrophil chemotaxis (n = 3 pooled samples from 9 mice). Accompanying heatmap displays representative downregulated genes associated with chemotaxis/migration and pro-inflammation. c, Flow cytometry analysis of cardiac neutrophils across treatment groups (Sham, I/R, VecDC, iCDC), showing changes in neutrophil proportion and its expression of ICAM-1 and CD14. iCDC treatment reduces the frequency of CD14+ and ICAM-1+ neutrophils.  d, Dot plot showing expression of representative marker genes across identified cardiac B cell subsets. e, GO and KEGG enrichment analysis of downregulated genes in B cells from iCDC-treated hearts versus I/R controls, highlighting suppression of B cell activation, immunoglobulin production, and ERK signalling pathways (n = 3 pooled samples from 9 mice). f, Flow cytometry analysis of cardiac B cells across treatment groups (Sham, I/R, VecDC, iCDC). Representative plots show decreased expression of activation markers CD69, CD80, and IgG in B cells from iCDC-treated hearts, indicating suppressed B cell activation. P values in b and e were calculated using a hypergeometric test.
Extended Data Fig. 11 iCDC treatment modulates cardiac fibroblast profiles in the ischaemia/reperfusion-injured mouse heart.
a, UMAP visualization of 139,036 single cells isolated from murine hearts post-I/R, clustered into 18 major cell types including fibroblasts, endothelial cells, macrophages, monocytes, dendritic cells, neutrophils, T cells, B cells, and natural killer (NK) cells. b, Dot plot showing expression of defining marker genes for fibroblast subclusters. c, GO and KEGG enrichment of downregulated genes in cardiac fibroblasts from iCDC-treated versus I/R control hearts, including pathways related to extracellular matrix (ECM) organization, cell-substrate adhesion, and wound healing (n = 3 pooled samples from 9 mice). d, Heatmap showing downregulated fibroblast genes involved in profibrotic and matrix-remodelling functions in fibroblasts from iCDC-treated hearts.   e, ECM regulator score of F-Myo fibroblasts across Sham, I/R, VecDC, and iCDC-treated groups shows significant downregulation in the iCDC group. n = 3 pooled samples from 9 animals. e, ECM regulator score of F-Myo fibroblasts across Sham, I/R, VecDC, and iCDC-treated groups shows significant downregulation in the iCDC group. n = 3 pooled samples from 9 animals.
f, ECM regulator scores across different subtypes of cardiac fibroblasts. F-Myo n = 7461, F-Act n = 7381, F-SH n = 18467, F-IFNs n = 922, F-SL n = 37945, F-IR n = 719 fibroblasts. g, Flow cytometry analysis of PDGFRα+ fibroblasts and their MHC-II expression in heart tissue reveals a reduced proportion of total fibroblasts and MHC-II+ fibroblasts in iCDC-treated hearts compared to I/R and VecDC groups. h, Quantitative RT-PCR analysis of Fibronectin, Periostin, and Acta2 expression in fibroblasts co-cultured with WTDC, VecDC, or iCDC (n = 3 biological replicates). Data are presented as mean ± s.e.m; all statistical tests are two-sided; Wilcoxon test (e, f), hypergeometric test (c), and one-way ANOVA with Tukey’s multiple-comparisons test (h).
Source data
Extended Data Fig. 12 Characterization and therapeutic evaluation of iCDCs in non-human primates.
a, Flow cytometry analysis of cultured CD11c+ dendritic cells in wild-type DCs (WTDC), vector control DCs (VecDC), and FAP-targeted iCDCs, showing the expression of co-stimulatory molecules (CD80, CD86, MHC-II). b, Quantification of end-systolic volume (ESV) and end-diastolic volume (EDV) by MRI at baseline baseline (D0), day 2 (D2), 1 month (M1), and 3 months (M3). n = 7 primates in MI, n = 10 primates in iCDC. Data are presented as mean ± s.e.m.
Source data
Extended Data Fig. 13 Systemic safety assessment of iCDCs therapy in non-human primates.
a, Haematological parameters-including white blood cell count (WBC), platelet count (PLT), and haemoglobin levels-as well as liver and kidney function markers, including albumin/globulin (A/G) ratio, alanine aminotransferase (ALT), aspartate aminotransferase (AST), and creatinine (Cr), were assessed at baseline (Day 0), and at 2, 4, 7, and 14 days, as well as 1, 2, and 3 months post-MI in both MI and iCDC-treated groups. No significant differences in WBC, PLT, haemoglobin levels, A/G, ALT, AST, and Cr were observed between the groups over time. n = 7 primates in MI, n = 10 primates in iCDC.  b, Circulating cytokine levels of IL-6, IL-10, IL-17A, and IFN-γ were assessed pre-MI, post-MI (prior to treatment), and at 4, 7, and 14 days post-MI. n = 7 primates in MI, n = 10 primates in iCDC. c, Representative hematoxylin and eosin (H&E) staining of major organs including liver, lung, colon, intestine, kidney, spleen, brain, skin, and thyroid collected at 3 months post-MI in MI and iCDC-treated animals. No overt histopathological abnormalities were observed in the iCDC group. d, Representative brain magnetic resonance imaging (MRI) scans from MI and iCDC-treated animals at 3 months post-MI, showing comparable brain morphology between groups. e, Quantification of non-sustained VT episodes and durations, and PVC counts in each non-human primate treated with or without iCDC daily during the 28-day monitoring. Data are presented as mean ± s.e.m., except for WBC, PLT, AST and Cr in a and b shown as mean ± s.d.
Source data
Extended Data Fig. 14 Gating strategies used for flow cytometry analysis and cell sorting.
a, Gating strategy for analysis of co-stimulatory molecules (CD80, CD86, MHC-II), and inhibitory checkpoints (PD-L1, TIM3) in bone marrow-derived CD11c+ dendritic cells. b, Gating strategy for quantification of T cell activation (CD69+), and classification of T cell subsets: naive T cells (CD44−CD62L+), effector memory T cells (TEM, CD44+CD62L−), and central memory T cells (TCM, CD44+CD62L+). c–d, Gating strategy for identification of CD4+ T cells, CD8+ T cells, TH17 cells (CD3+CD4+IL-17+), and regulatory T cells (Tregs, CD3+CD4+CD25+Foxp3+).   e, Gating strategy for delineation of different myeloid cell subsets: neutrophils (CD45+CD11b+Ly6G+), CCR2+ macrophages, CCR2−TIM4+ macrophages, and CCR2− MHC-II+ macrophages. e, Gating strategy for delineation of different myeloid cell subsets: neutrophils (CD45+CD11b+Ly6G+), CCR2+ macrophages, CCR2−TIM4+ macrophages, and CCR2− MHC-II+ macrophages.
f, Gating strategy for evaluation of ICAM-1+ neutrophils, CD14+ neutrophils, and IgG+ B cells, IgM+ B cells in the mouse hearts. g, Gating strategy for analysis of fibroblasts (CD45−CD140α+) and activated fibroblast subsets expressing MHC class II.
Supplementary informationSource dataRights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Cite this article
Li, X., Li, J., Li, G. et al. Engineered immunosuppressive dendritic cells protect against cardiac remodelling.
Nature (2026). https://doi.org/10.1038/s41586-026-10346-5
Download citation
Received: 23 June 2025
Accepted: 02 March 2026
Published: 08 April 2026
Version of record: 08 April 2026
DOI: https://doi.org/10.1038/s41586-026-10346-5 |









